Introduction
I only recently discovered the fundamental connection between the C and R languages. It was during a Bay Area useR Group meeting, where presenter J.J. Allaire shared two points to motivate his talk on Rcpp. The first explained just how much of modern R really is C and C++. For illustration, he used the librestats language composition analysis of Core R, which might not have been so interesting if it did not extend to R's contributed packages. Allaire's second point emphasized R's roots as an interactive mechanism for calling code from compiled languages like C and Fortran (which you can also hear from S inventor John Chambers). While I didn't understand all of the finer points at the time, I never thought of R in the same way again.
Wanting to learn more about this connection between languages, I dug into the C calling paradigms of R. My own lack of experience was mitigated by adopting a tremendously limited scope: the function f(x)= 2x. In the course of its implementation, I received advice from respected voices, each highlighting important points about the alternatives. In this article, I'll share what I learned as concisely as possible. So if you have a passing familiarity with C and an interest in R, grab a cup of coffee, pop open a terminal, and prepare to explore .C, .Call, and Rcpp's sourceCpp.
The big three
The .C function interface
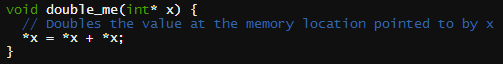
Of R's native functions, the .C interface is the simplest but also the most limited way to call C from R. Inside a running R session, the .C interface allows objects to be directly accessed in an R session's active memory. Thus, to write a compatible C function, all arguments must be pointers. No matter the nature of your function's return value, it too must be handled using pointers. The C function you will write is effectively a subroutine.Our function f(x)= 2x, implemented as double_me in the file doubler.c, is shown below.
void double_me(int* x) {
// Doubles the value at the memory location pointed to by x
*x = *x + *x;
}
To compile the C code, run the following line at your terminal:
$ R CMD SHLIB doubler.cIn an R interactive session, run:
dyn.load("doubler.so")
.C("double_me", x = as.integer(5))
$x [1] 10Notice that the output of .C is a list with names corresponding to the arguments. While the above code is pure C, adding C++ code (instead of C) is made possible by using the extern wrapper.
.Call
The .Call interface is the more fully featured and complex cousin of the .C interface. Unlike .C, .Call requires header files that come standard with every R installation. These header files provide access to a new data type, SEXP. The following code, stored in the file, doubler2.c, illustrates its use.
#include <R.h>
#include <Rdefines.h>
SEXP double_me2(SEXP x) {
// Doubles the value of the first integer element of the SEXP input
SEXP result;
PROTECT(result = NEW_INTEGER(1)); // i.e., a scalar quantity
INTEGER(result)[0] = INTEGER(x)[0] * 2;
UNPROTECT(1); // Release the one item that was protected
return result;
}
Unlike our experience with the .C interface, double_me2 is a function and does return a value. While that appeals to intuition, no matter what the native input and output types, they must now live in a SEXP object. To code double_me2, you must know that there's an integer in the input x, and extract it as if it were the first item in a C array. For the return value, you must add your integer result to a SEXP object in an equally unnatural way. The PROTECT function must be used to prevent R's automatic garbage collection from destroying all the objects.
As before, use R at the command line to compile doubler2.c:
$ R CMD SHLIB doubler2.cBack in the R interactive console, the steps are very similar.
dyn.load("doubler2.so")
.Call("double_me2", as.integer(5))
[1] 10Notice now that the output is an integer vector instead of a list.
Rcpp and the sourceCpp function
The .C and .Call examples above owe a debt to Jonathan Callahan's entries 8 and 10 of his Using R series. When the examples started working, I tweeted to share my excitement. An hour later, I saw a familiar face:
Let's check it out.
In terms of the code alone, it's easy to see where Hadley is coming from. It's readable, looks just like standard C++ code, and features data types that make intuitive sense. Our simple function is implemented below, saved in the final static file doubler3.cpp (though, in all humility, it's really just C).
#include <Rcpp.h>
using namespace Rcpp;
// [[Rcpp::export]]
int double_me3(int x) {
// takes a numeric input and doubles it
return 2 * x;
}
I'll refer you to Hadley's article High performance functions with Rcpp for details on Rcpp, but for now, note the "// [[Rcpp::export]]" comment, necessary before each C/C++ function, and the updated #include statement. Most importantly, notice how the pointers and SEXP objects have been replaced. Just like our original function f(x), double_me3 takes one integer input and returns one integer output.
After installing the Rcpp package, we're back to the console one final time.
library(Rcpp)
sourceCpp("doubler3.cpp")
double_me3(5)
[1] 10With Rcpp, the function is waiting for us in the global environment, without even compiling at the command line. Pretty convenient!
Discussion
With the disclaimer that I am a C++ novice, I summarize my thoughts on the three options below.
I like the simplicity of the C code written for the .C interface. It doesn't rely on external header files and it is possible to test using a C compiler alone. On the other hand, I don't like that a function has to be morphed into a subroutine that uses pointers.
In the .Call interface, SEXP objects are also pointers, though that is perhaps superfluous. My biggest complaint is the verboseness that was added to our short example. As Jonathan Callahan points out, .Call "requires much more knowledge of R internals [but] is the recommended, modern approach for serious C [and C++] programmers."
After seeing Rccp in action, it's not hard to understand why Hadley sent me directly to Rcpp. The C code looks great, there were fewer steps, and our function was ready for us inside the R global environment.
Perhaps you're wondering if there is any reason not to use Rcpp. According Murray Stokely, Google Software Engineer, it could be risky on a very large project. "Rcpp's heavy reliance on C macros can make it unsafe to use with large code bases," says Stokely. For example, the Rcpp FAQ (Section 3.9) describes an unresolved issue casting 64-bit integer types. The implication, Stokely explains, is that a loss of precision could occur without any errors or warnings. The Rcpp FAQ considers such examples "corner cases," and perhaps the typical user will not have to worry.
Whatever your decision, I wish you the best on your C++ journey!





